(Re)Discovering Protein Structure and Function Through Language Modeling (ArXiv)(Code)
- In our study, we show how a language model, trained simply to predict a masked (hidden) amino acid in a protein sequence, recovers high-level structural and functional properties of proteins. In particular, we show how the Transformer language model uses attention (1) to capture the folding structure of proteins, connecting regions that are apart in the underlying sequence but spatially close in the protein structure, and (2) targets binding sites, a key functional component of proteins. We also introduce a three-dimensional visualization of the interaction between attention and protein structure. Our findings align with biological processes and provide a tool to aid scientific discovery. The code for the visualization tool and experiments is available at https://github.com/salesforce/provis.
- TL;DR: Trained solely on language modeling, the Transformer’s attention mechanism recovers high-level structural and functional properties of proteins.
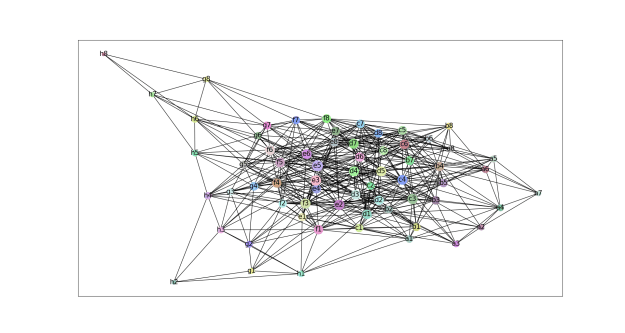
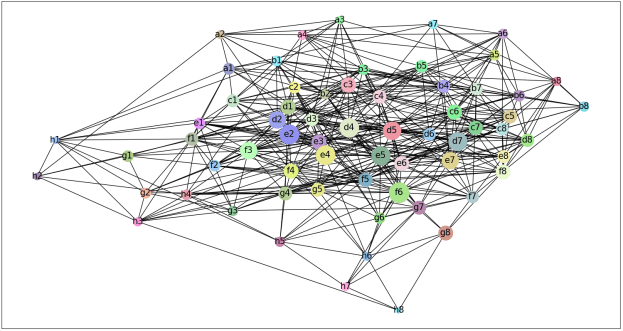
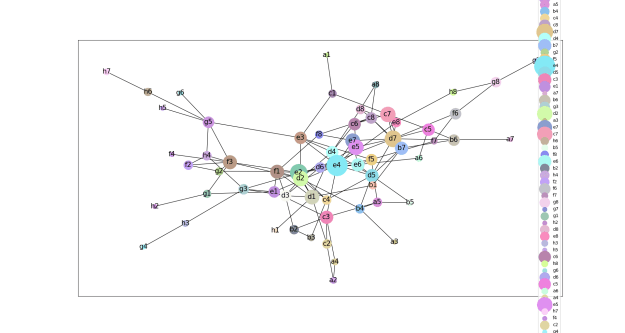
- We explored the degree to which attention captures these contact relationships by analyzing the attention patterns of 5,000 protein sequences and comparing them to ground-truth contact maps. Our analysis revealed that one particular head — the 12th layer’s 4th head, denoted as head 12-4 — aligned remarkably well with the contact map. For “high confidence” attention (> .9 ), 76% of this head’s total attention connected amino acids that were in contact. In contrast, the background frequency of contacts among all amino acid pairs in the dataset is just 1.3%.
GPT-2 Agents
- Add a menu that writes node spatial information to the DB
- Add a “Graph from DB” menu that assembles the edge information from the move table and the node information from the new table, above.
- Continue on path finding
- Distance between a point and a line using numpy (stackoverflow). Not exactly what I need, which is the point of intersection and the distance. There is a stackoverflow post that is close, but here’s a version that tests the results and plots it:
import numpy as np import math import matplotlib.pyplot as plt p1 = np.array([1.0, 1.0]) l1 = np.array([0.0, 1.0]) l2 = np.array([1.0, 0.0]) lvec = l2 - l1 lvec /= np.linalg.norm(lvec, 2) p2 = l1 + lvec * np.dot(p1 - l1, lvec) print("intesection = {}".format(p2)) #0.2 1. pvec = p2 - p1 dist = np.linalg.norm(pvec, 2) pvec /= dist det = np.linalg.det([lvec, pvec]) dot = np.dot(lvec, pvec) rads = math.atan2(det, dot) print("distance = {}, angle = {}".format(dist, math.degrees(rads))) plt.plot([l1[0], l2[0]],[l1[1], l2[1]]) plt.plot([p1[0], p2[0]],[p1[1], p2[1]]) plt.show()
- Distance between a point and a line using numpy (stackoverflow). Not exactly what I need, which is the point of intersection and the distance. There is a stackoverflow post that is close, but here’s a version that tests the results and plots it:
- Here’s the test for seeing if a point is on a line. Again, loosely based on a stackoverflow post:
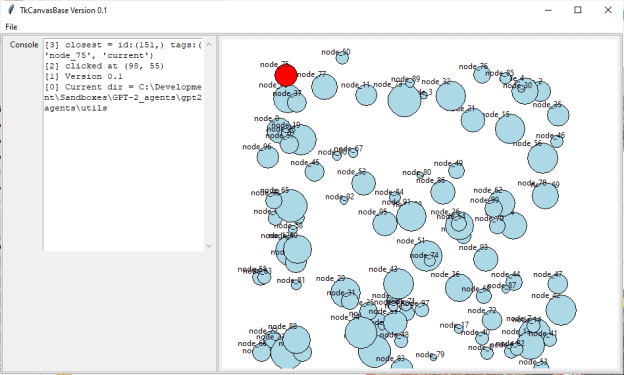
def is_between(self, l1:[int, int], l2:[int, int], p1:[int, int], epsilon:float = .1) -> bool: p1 = np.array(p1).astype(np.float) l1 = np.array(l1).astype(np.float) l2 = np.array(l2).astype(np.float) s1 = np.linalg.norm(l1-p1) s2 = np.linalg.norm(l2-p1) d = np.linalg.norm(l2-l1) # print("d = {}, s1 + s2 = {}".format(d, s1+s2)) if abs(d - (s1+s2)) < epsilon: return True return False - Got graphical node selection working. Need to tie that back into the menus for start and stop
Proposal
- Looks like no writing today. Done, maybe?
GOES
- 10:00 CASSIE demo – really good
- 12:00 All Hands – need to catch up on my training. Something for the afternoons?
ML Seminar
- Status report
- Participated in some some triage on Arpita’s and Fatima’s paper































You must be logged in to post a comment.