7:00 – 8:00 Research
- Modeling the Evolution of Ideological Landscapes Through Opinion Dynamics
- More poster. Tightening *everything*. Progress! Should be ready for Don on Thursday
8:30 – 3:00 BRC
- Switching gears from LaTex to Python takes effort. Neither is natural or comfortable yet
- Sent Jeremy a note on conferences and vacation. Using the hours on my paycheck stub, which *could* be correct…
- More clustering. Adding output that will be used for the optimizer
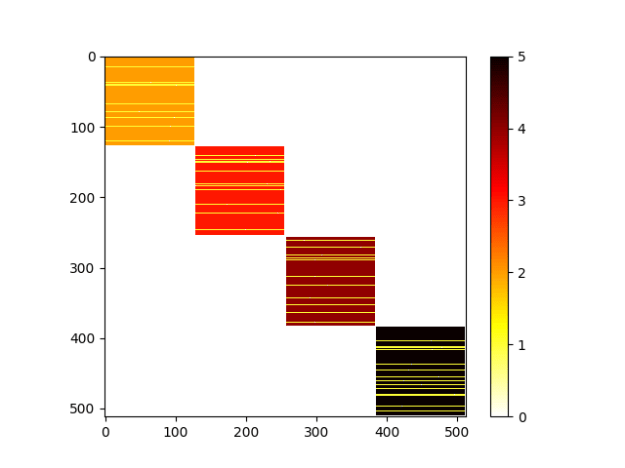
clusters = 4 Total = 512 clustered = 437 unclustered = 75
- Built out the optimizer and filled it with a placeholder function. Will fill in after lunch
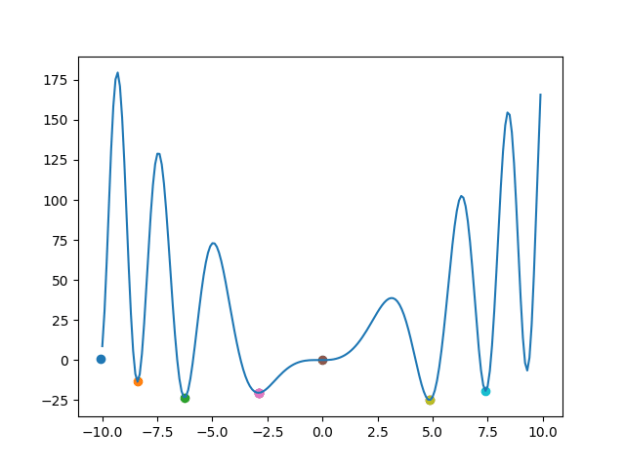
- Had to leave to take care of dad, who fainted. But here are my thoughts on the GA construction. The issue with fitness test is that we have two variables to optimize, the EPS and the minimum cluster size, based on the number of clusters and the number of unclustered. I want to unitize the outputs sop that 2.0 is best and 0.0 is worst. The unclustered should be 1.0 – unclustered/total. The number of clusters should be clusters/(total/min_cluster_size).
- The way the GA should work is that we start with a set of initial EPSs (0 – 1) and a set of cluster sizes (3 – total/3). We try each, throw the bottom half away, keep the top result and breed a new set by interpolating (random distances?) between the remaining. We also randomly generate a new allele or two in case we get trapped on a local maxima. When we are no longer getting any improvement (some epsilon) we stop. All the points can be plotted and we can try to fit a polyline as well (one for eps and for minimum cluster? Could plot as a surface…)
