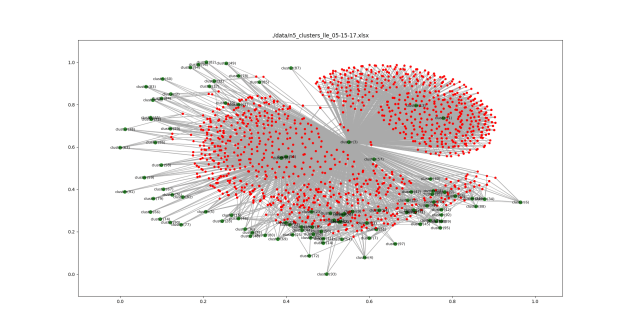
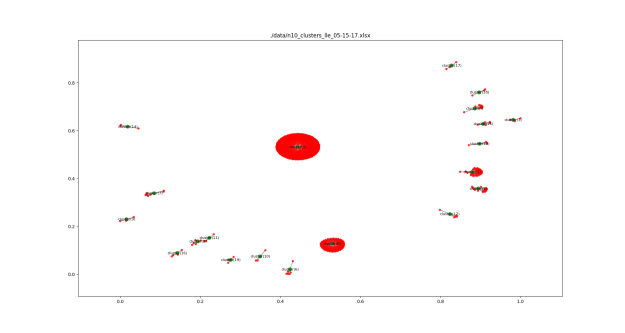
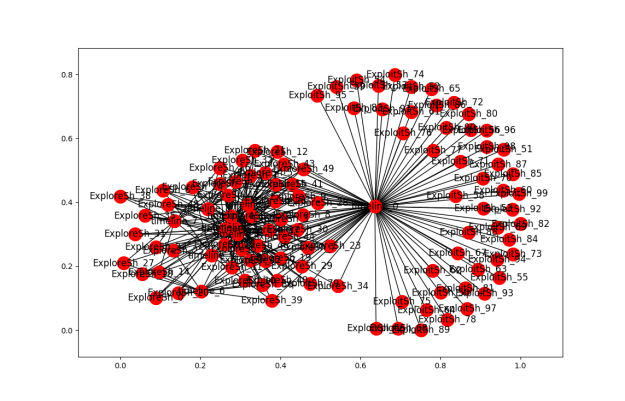
More clustering. Here’s the list of agents by clusters. An OPEN state means that the simulation finished with agents in the cluster. Num_entries is: the lifetime of the cluster. For these runs, the max is 200. Id is the ‘name’ of the cluster. Tomorrow, I’ll try to get this drawn using networkx.
timeline[0]:
Id = cluster_0
State = ClusterState.OPEN
Num entries = 200
{'ExploitSh_52', 'ExploreSh_43', 'ExploitSh_56', 'ExploreSh_2', 'ExploreSh_5', 'ExploitSh_73', 'ExploitSh_95', 'ExploreSh_19', 'ExploreSh_4', 'ExploitSh_87', 'ExploitSh_76', 'ExploreSh_3', 'ExploitSh_93', 'ExploreSh_32', 'ExploreSh_41', 'ExploreSh_17', 'ExploitSh_88', 'ExploitSh_77', 'ExploreSh_39', 'ExploitSh_85', 'ExploreSh_40', 'ExploitSh_64', 'ExploreSh_34', 'ExploreSh_22', 'ExploitSh_99', 'ExploreSh_1', 'ExploitSh_97', 'ExploitSh_69', 'ExploreSh_29', 'ExploitSh_58', 'ExploitSh_62', 'ExploreSh_23', 'ExploreSh_36', 'ExploreSh_11', 'ExploitSh_80', 'ExploitSh_82', 'ExploreSh_21', 'ExploitSh_75', 'ExploitSh_72', 'ExploitSh_89', 'ExploitSh_86', 'ExploreSh_37', 'ExploitSh_84', 'ExploitSh_81', 'ExploreSh_15', 'ExploitSh_51', 'ExploreSh_44', 'ExploitSh_83', 'ExploitSh_94', 'ExploreSh_16', 'ExploitSh_53', 'ExploitSh_67', 'ExploitSh_74', 'ExploreSh_45', 'ExploreSh_26', 'ExploreSh_12', 'ExploreSh_13', 'ExploitSh_92', 'ExploreSh_9', 'ExploreSh_28', 'ExploitSh_50', 'ExploreSh_8', 'ExploreSh_30', 'ExploreSh_49', 'ExploitSh_59', 'ExploitSh_57', 'ExploreSh_42', 'ExploitSh_65', 'ExploitSh_54', 'ExploitSh_61', 'ExploitSh_66', 'ExploitSh_55', 'ExploitSh_78', 'ExploitSh_68', 'ExploitSh_79', 'ExploitSh_91', 'ExploitSh_71', 'ExploreSh_7', 'ExploitSh_98', 'ExploitSh_60', 'ExploitSh_70', 'ExploreSh_10', 'ExploitSh_90', 'ExploreSh_46', 'ExploitSh_96', 'ExploreSh_47', 'ExploitSh_63'}
timeline[1]:
Id = cluster_1
State = ClusterState.OPEN
Num entries = 200
{'ExploreSh_25', 'ExploreSh_6', 'ExploreSh_38', 'ExploreSh_43', 'ExploreSh_49', 'ExploreSh_1', 'ExploreSh_2', 'ExploreSh_20', 'ExploreSh_33', 'ExploreSh_48', 'ExploreSh_5', 'ExploreSh_29', 'ExploreSh_15', 'ExploreSh_42', 'ExploreSh_24', 'ExploreSh_19', 'ExploreSh_4', 'ExploreSh_44', 'ExploreSh_16', 'ExploreSh_23', 'ExploreSh_36', 'ExploreSh_11', 'ExploreSh_3', 'ExploreSh_27', 'ExploreSh_35', 'ExploreSh_32', 'ExploreSh_17', 'ExploreSh_26', 'ExploreSh_21', 'ExploreSh_12', 'ExploreSh_18', 'ExploreSh_45', 'ExploreSh_41', 'ExploitSh_79', 'ExploreSh_13', 'ExploreSh_0', 'ExploreSh_39', 'ExploreSh_7', 'ExploreSh_9', 'ExploreSh_28', 'ExploreSh_40', 'ExploreSh_31', 'ExploreSh_10', 'ExploreSh_46', 'ExploreSh_37', 'ExploreSh_14', 'ExploreSh_47', 'ExploreSh_8', 'ExploreSh_30', 'ExploreSh_34', 'ExploreSh_22'}
timeline[2]:
Id = cluster_2
State = ClusterState.CLOSED
Num entries = 56
{'ExploreSh_25', 'ExploreSh_1', 'ExploreSh_33', 'ExploreSh_29', 'ExploreSh_5', 'ExploreSh_48', 'ExploreSh_15', 'ExploreSh_19', 'ExploreSh_36', 'ExploreSh_3', 'ExploreSh_11', 'ExploreSh_35', 'ExploreSh_45', 'ExploreSh_17', 'ExploreSh_26', 'ExploreSh_41', 'ExploitSh_79', 'ExploreSh_13', 'ExploreSh_9', 'ExploreSh_40', 'ExploreSh_31', 'ExploreSh_37', 'ExploreSh_47', 'ExploreSh_30', 'ExploreSh_22'}
timeline[3]:
Id = cluster_3
State = ClusterState.CLOSED
Num entries = 16
{'ExploreSh_25', 'ExploreSh_6', 'ExploreSh_43', 'ExploreSh_2', 'ExploreSh_48', 'ExploreSh_5', 'ExploreSh_15', 'ExploreSh_42', 'ExploreSh_24', 'ExploreSh_4', 'ExploreSh_44', 'ExploreSh_3', 'ExploreSh_26', 'ExploreSh_17', 'ExploreSh_41', 'ExploreSh_21', 'ExploreSh_32', 'ExploreSh_13', 'ExploreSh_9', 'ExploreSh_7', 'ExploreSh_28', 'ExploreSh_37', 'ExploreSh_8', 'ExploreSh_30', 'ExploreSh_49', 'ExploreSh_22'}
timeline[4]:
Id = cluster_4
State = ClusterState.CLOSED
Num entries = 30
{'ExploreSh_6', 'ExploreSh_1', 'ExploreSh_2', 'ExploreSh_20', 'ExploreSh_33', 'ExploreSh_48', 'ExploreSh_15', 'ExploreSh_24', 'ExploreSh_4', 'ExploreSh_16', 'ExploreSh_23', 'ExploreSh_3', 'ExploreSh_11', 'ExploreSh_26', 'ExploreSh_41', 'ExploreSh_17', 'ExploreSh_32', 'ExploreSh_18', 'ExploreSh_13', 'ExploreSh_9', 'ExploreSh_46', 'ExploreSh_37', 'ExploreSh_8', 'ExploreSh_30', 'ExploreSh_49', 'ExploreSh_22'}
timeline[5]:
Id = cluster_5
State = ClusterState.CLOSED
Num entries = 28
{'ExploreSh_25', 'ExploreSh_43', 'ExploreSh_2', 'ExploreSh_48', 'ExploreSh_29', 'ExploreSh_42', 'ExploreSh_24', 'ExploreSh_4', 'ExploreSh_44', 'ExploreSh_36', 'ExploreSh_35', 'ExploreSh_45', 'ExploreSh_17', 'ExploreSh_26', 'ExploreSh_12', 'ExploreSh_0', 'ExploreSh_28', 'ExploreSh_40', 'ExploreSh_31', 'ExploreSh_46', 'ExploreSh_37', 'ExploreSh_14', 'ExploreSh_47', 'ExploreSh_8', 'ExploreSh_30', 'ExploreSh_22'}
timeline[6]:
Id = cluster_6
State = ClusterState.CLOSED
Num entries = 10
{'ExploreSh_40', 'ExploreSh_25', 'ExploreSh_18', 'ExploreSh_27', 'ExploreSh_10', 'ExploreSh_13', 'ExploreSh_20', 'ExploreSh_0', 'ExploreSh_37', 'ExploreSh_14', 'ExploreSh_36', 'ExploreSh_11', 'ExploreSh_39', 'ExploreSh_42', 'ExploreSh_22'}
timeline[7]:
Id = cluster_7
State = ClusterState.CLOSED
Num entries = 9
{'ExploreSh_38', 'ExploreSh_2', 'ExploreSh_4', 'ExploreSh_46', 'ExploreSh_16', 'ExploreSh_33', 'ExploreSh_47', 'ExploreSh_14', 'ExploreSh_11', 'ExploreSh_27', 'ExploreSh_35', 'ExploreSh_45'}
timeline[8]:
Id = cluster_8
State = ClusterState.CLOSED
Num entries = 25
{'ExploreSh_21', 'ExploreSh_38', 'ExploreSh_19', 'ExploreSh_2', 'ExploreSh_13', 'ExploreSh_44', 'ExploreSh_1', 'ExploreSh_10', 'ExploreSh_16', 'ExploreSh_47', 'ExploreSh_5', 'ExploreSh_48', 'ExploreSh_42', 'ExploreSh_35', 'ExploreSh_22', 'ExploreSh_32'}
timeline[9]:
Id = cluster_9
State = ClusterState.OPEN
Num entries = 16
{'ExploreSh_17', 'ExploreSh_6', 'ExploreSh_24', 'ExploreSh_19', 'ExploreSh_10', 'ExploreSh_20', 'ExploreSh_46', 'ExploreSh_33', 'ExploreSh_14', 'ExploreSh_3', 'ExploreSh_39', 'ExploreSh_7', 'ExploreSh_45'}
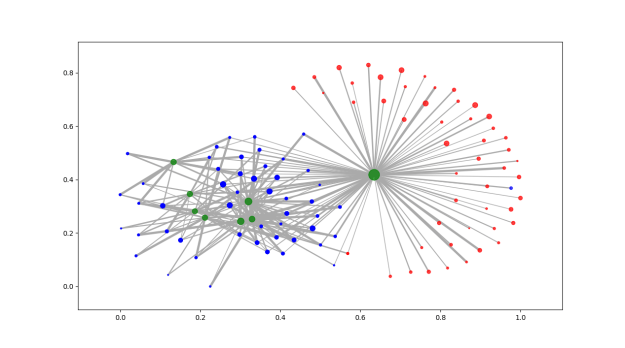
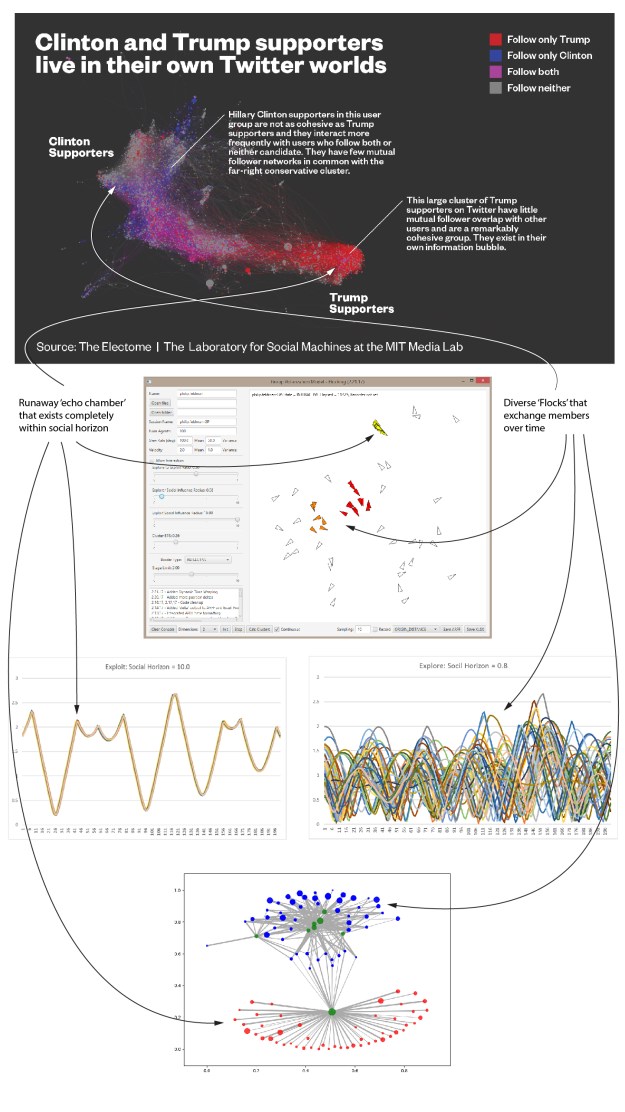
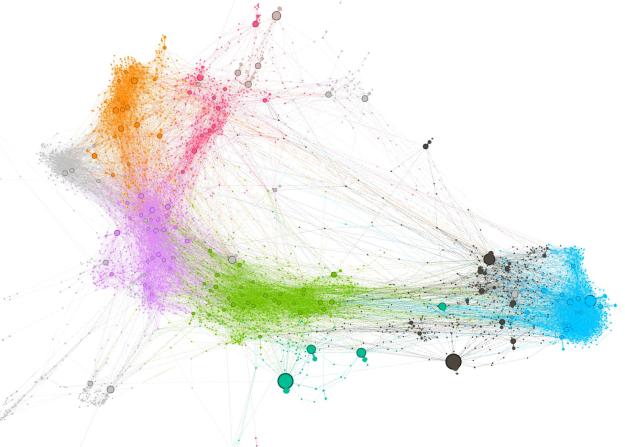
 Add a picture of a network next to each item that reflects the polarization level. Maybe gray out the non-abstract bits.
Add a picture of a network next to each item that reflects the polarization level. Maybe gray out the non-abstract bits.







You must be logged in to post a comment.